updateDetector
Update settings of a trained deep learning anomaly detector and recompute detection threshold
Since R2025a
Syntax
Description
Add-On Required: This feature requires the Time Series Anomaly Detection for MATLAB add-on.
Set New Threshold Manually
detector = updateDetector(detector,ThresholdMethod="manual",Threshold=Threshold)Threshold that you
specify. Use this syntax when you want to adjust the threshold directly rather than
recompute it from input time series data.
For examples that you can open and experiment with, see Use UpdateDetector to Tune Anomaly Detector Performance Without Retraining
This function requires Deep Learning Toolbox™.
Recompute Threshold Using New Data
detector = updateDetector(detector,data)detector using the
data in data and the current settings of the detector model, as long
as detector.Threshold is not set to
"manual".
Use this syntax when you want to compute the threshold using a different set of data
than the original training data. The data data that you use must
consist only of only normal data, that is, that contains no known anomalies or anomalous
data.
Set Additional Options
detector = updateDetector(___,Name=Value)Name=Value arguments. These
options set the corresponding properties of detector, include
thresholding and execution-related properties. The options you can set depend on the
detector model you are updating.
Note that, when you use updateDetector, there are no default
values for model settings, as the model is already configured. You must explicitly specify
any settings, such as the threshold method and the corresponding threshold parameter, that
differ from the initial detector settings.
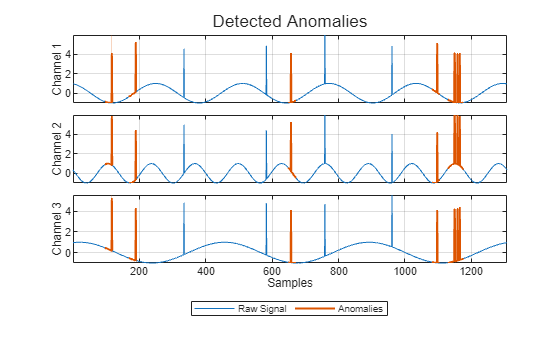
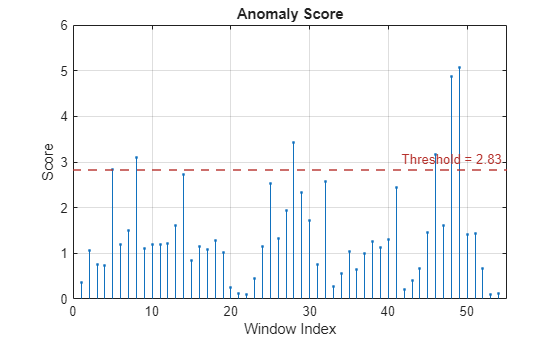
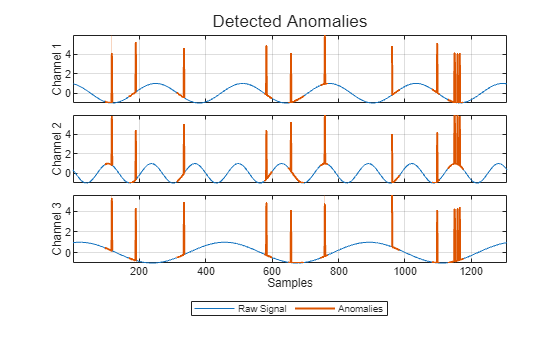
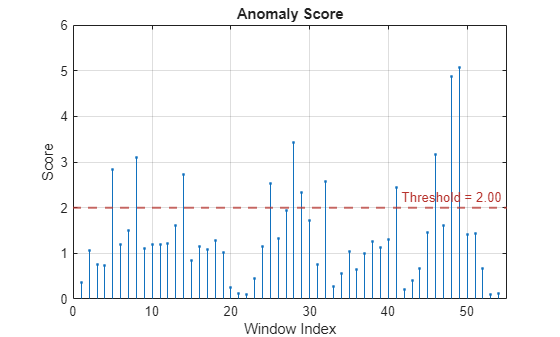
For example, suppose you first create a time series isolated forest detector model
using the "kSigma" threshold method, which has an associated default
threshold parameter of 3. After viewing the results, you decide to try
the "ContaminationFraction" method instead, using a threshold parameter
value that identifies the top 5% of the anomaly scores as anomalous. To do so, use the
following command. detector =
updateDetector(detector,data,ThresholdMethod="contaminationFraction",ThresholdParameter=0.05)