Simulate Groups Using Doses and Variants from Data Set
The Group Simulation program in the SimBiology Model Analyzer app lets you simulate various groups, such as patients, from your data set. If the data set also has dosing information and group-specific variants, the program lets you apply them to each group during simulation. It also lets you reuse the same program setup and perform parameter estimation. The following steps illustrate how to simulate the tumor weight of various patients and reuse the same program configuration to estimate some model parameters.
Load Tumor Growth Model and Data
Enter the following command to load the tumor growth model and data in SimBiology Model Analyzer.
openExample('simbio/GroupSimulationSimBiologyModelAnalyzerExample')
In the Browser pane (if it is in the default Project View), expand the Data1 folder and double-click Datasheet1 to open the data set.

If your browser is in the Workspace View, click Documents and double-click Datasheet1 instead.

The app automatically classifies the following columns:
The ID column as group (a grouping variable).
The Time column as independent (an independent variable).
The Dose column as dose1 (a dosing variable). If the data has more than one dose columns, you can classify them as dose2, dose3, and so on.
For the measured response data columns such as CentralConc, you need to manually classify them as the dependent variables. Double-click Classification under CentralConc and select dependent. A dependent variable is a response that is time-varying.
Repeat the same process for PeripheralConc and TumorWeight.
Tip: The data set also contains two variant columns k1 and Cl_Central, which contain group-specific values for the parameters k1 and Cl_Central. Optionally, you can classify them as covariate. A covariate is a value that a response may depend on and could be used as a category variable in plotting, for instance. For details on setting up group-specific variant columns, see variableNames.

Set up Group Simulation Program Using Doses and Variants from Data Set
Select Program > Group Simulation to open a group simulation program. The Model and Data steps of the program have been prepopulated with Tumor Growth Model and Data1, respectively.
In the Data Map table, clear CentralConc and PeripheralConc. Keep TumorWeight selected.
Double-click the Component cell next to TumorWeight and enter tumor. Select [Tumor Growth Model].tumor_weight from the drop down menu.

Apply the doses from the data set. In the dose from data row, map the Dose column to Central.Drug to indicate that the Drug species in the Central compartment is being dosed.

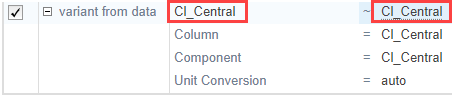
To apply group-specific variants from the data set, click the variant button ![]() . A variant from data row appears. Double-click the cell named Column and enter the name of the variant column from the data set to use: k1. In the next cell, enter the corresponding component name, which is also k1.
. A variant from data row appears. Double-click the cell named Column and enter the name of the variant column from the data set to use: k1. In the next cell, enter the corresponding component name, which is also k1.

Similarly, map the Cl_Central column.
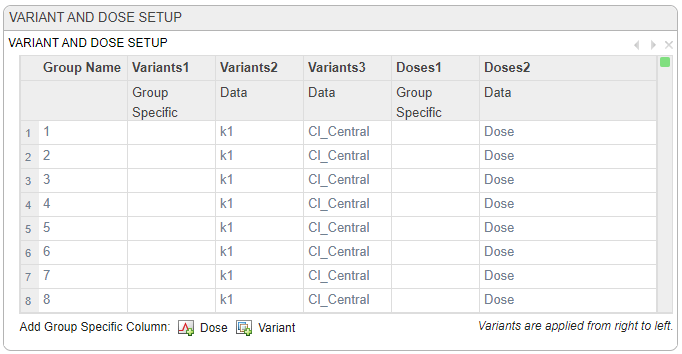
The Variant and Dose Setup table is automatically updated to indicate that the variants and doses from the data are now being applied to the model. If all values of a variable within a group are NaN, the program creates no entry in the variant content for the group. When you simulate the model with the variant, the program uses the associated model component value as the initial value for model simulation.
Run Group Simulation
In the Simulation step, you can select whether to include the time points from the data set only, include solver time points only, or include both during model simulation. You must select at least one option.
Time points contain data times within each group — Include the exact time points for each group from data, such as the times listed in the Time column of the input data set.
Time points contain solver time points — Include time points used by the solver during model simulation. This option lets you define a custom stop time or pick a maximum stop time from data. There are three choices to pick from.
a. Simulate to maximum time within each group — Use the group-specific maximum time. Each group simulates to its own stop time.
b. Simulate to maximum time across all groups — Use a single stop time across all the groups. The app checks all the stop times from the groups and picks the maximum.
c. Stop Time — Use a custom stop time. If the last time point of a group from the data set is greater than the specified stop time, the program ignores the stop time and simulates until that last time point for that group.
You can also add sliders to vary model quantity values in the Explorer panel, and the Slider Summary section displays each of those quantities. For now, leave it empty.
To check other simulation settings, such as solver type and absolute tolerance, point the mouse to the top right and click the option menu button. For the purposes of this example, use the default simulation options that are already selected as shown in the following figure.

Click the Run button of the step to simulate the groups.
The program opens the Plot1 tab to show the simulated tumor weight response for each group as shown next. The solid lines are the simulation results while the dotted lines with circle markers represent the experimental data.

Estimate Parameters Using Same Setup as Group Simulation
The program lets you fit model parameters to data using the same setup you use for group simulation without having to reconfigure or create a separate Fit Data program.
Click the Program1 tab. At the top of the Simulation step, select Simulation > Fit.

Click Yes if a dialog asks whether to continue. The step changes to the Fit step.
Tip: Switching steps here removes the results of the step that you just ran. To avoid the results from being removed, you can save the results first before switching. In the Browser panel, expand Program1. Right-click LastRun and select Save Data.
In the Estimated Parameters table, double-click the empty row and enter L0. Similarly, enter L1 as shown in the following figure.

At the bottom of the table, select Pooled Fit to estimate one set of parameter values for all groups.
![]()
Leave the rest of the settings as default. Note that the default estimation method is lsqnonlin if you have Optimization Toolbox™. If you do not, the app uses fminsearch.

Click the Run button of the Fit step to estimate parameters. The program next opens several tabs for the fit results and plots. For example, Datasheet2 contains the pooled parameter estimates and other fit statistics, such as AIC. For a more detailed fitting workflow example, see Calculate NCA Parameters and Fit Model to PK/PD Data Using SimBiology Model Analyzer.
Perform Group Simulation Using Parameter Estimates
You can switch back to the simulation step and simulate the groups using the pooled parameter estimates without applying the group-specific variants this time.
Click the Program1 tab. At the top of the Fit step, select Fit > Simulation. Click Yes if a dialog shows up to confirm. The step changes to Simulation.

In the Slider Summary section of the step, point the mouse to the top right, click the option menu button, and select Update Sliders to Match Fit Estimates.

The table now lists L0 and L1 parameters. The Value column contains the estimated values transferred from the previous Fit step.

Expand the Explorer panel. It now contains sliders for L0 and L1. The current value for each parameter is displayed on the top right of the slider bar as shown next.

In the Data Map table, clear the variant rows for k1 and Cl_Central.

Run the Simulation step. The program simulates the groups using the parameter estimates without applying group-specific variants and opens Plot1.

You can also explore the model behavior by moving the sliders in the Explorer panel to different parameter values or applying just some of the group-specific variants or doses.
Tip: Each time you move a slider, the program automatically simulates by default. You can change this default behavior by going to Preferences > Programs > Run program when slider changes value. Set the value to false. Then the program reruns only when you hit the Run button.
See Also
Topics
- Create Data File with SimBiology Definitions
- Calculate NCA Parameters and Fit Model to PK/PD Data Using SimBiology Model Analyzer
- Find Important Tumor Growth Parameters with Local Sensitivity Analysis Using SimBiology Model Analyzer
- Explore Biological Variability with Virtual Patients Using SimBiology Model Analyzer
- Scan Dosing Regimens Using SimBiology Model Analyzer App
- Scan Tumor Growth Using Group-Specific Doses and Variant Parameters from Data Set