LocalOutlierFactor
Description
Use a local outlier factor model object
LocalOutlierFactor for anomaly detection.
Outlier detection (detecting anomalies in training data) — Detect anomalies in training data by using the
loffunction. Theloffunction creates aLocalOutlierFactorobject and returns anomaly indicators and scores (local outlier factor values) for the training data.Novelty detection (detecting anomalies in new data with uncontaminated training data) — Create a
LocalOutlierFactorobject by passing uncontaminated training data (data with no outliers) tolof, and detect anomalies in new data by passing the object and the new data to the object functionisanomaly. Theisanomalyfunction returns anomaly indicators and scores for the new data.
Creation
Create a LocalOutlierFactor object by using the lof
function.
Properties
This property is read-only.
Predictors used to train the local outlier factor model, specified as a numeric
matrix or a table. Each row of X corresponds to one observation,
and each column corresponds to one variable.
This property is read-only.
Maximum number of data points in each leaf node of the Kd-tree, specified as a positive integer.
This property is valid when SearchMethod
is 'kdtree'. If SearchMethod is
'exhaustive', the BucketSize value is empty
([]).
This property is read-only.
Categorical predictor
indices, returned as a vector of positive integers. CategoricalPredictors
contains index values indicating that the corresponding predictors are categorical. The index
values are between 1 and p, where p is the number of
predictors used to train the model. If none of the predictors are categorical, then this
property is empty ([]).
This property is read-only.
Fraction of anomalies in the training data, specified as a numeric scalar in the
range [0,1].
If the
ContaminationFractionvalue is 0, thenloftreats all training observations as normal observations, and sets the score threshold (ScoreThresholdproperty value) to the maximum anomaly score value of the training data.If the
ContaminationFractionvalue is in the range (0,1], thenlofdetermines the threshold value (ScoreThresholdproperty value) so that the function detects the specified fraction of training observations as anomalies.
This property is read-only.
Distance metric, specified as a character vector.
If all the predictor variables are continuous (numeric) variables, then the
Distancevalue can be one of these distance metrics.Value Description 'euclidean'Euclidean distance
"fasteuclidean"Euclidean distance using an algorithm that usually saves time when the number of elements in a data point exceeds 10. See Algorithms.
"fasteuclidean"applies only to the"exhaustive"SearchMethod.'mahalanobis'Mahalanobis distance — The distance uses the covariance matrix stored in the
DistanceParameterproperty.'minkowski'Minkowski distance — The distance uses the exponent value stored in the
DistanceParameterproperty.'chebychev'Chebychev distance (maximum coordinate difference)
'cityblock'City block distance
'correlation'One minus the sample correlation between observations (treated as sequences of values)
'cosine'One minus the cosine of the included angle between observations (treated as vectors)
'spearman'One minus the sample Spearman's rank correlation between observations (treated as sequences of values)
If all the predictor variables are categorical variables, then the
Distancevalue can be one of these distance metrics.Value Description 'hamming'Hamming distance, which is the percentage of coordinates that differ
'jaccard'One minus the Jaccard coefficient, which is the percentage of nonzero coordinates that differ
For more information on the various distance metrics, see Distance Metrics.
This property is read-only.
Distance metric parameter value for the Mahalanobis or Minkowski distance, specified
as a positive scalar. The DistanceParameter value is empty
([]) for the other distances, indicating that the specified
distance metric formula has no parameters.
If
Distanceis'mahalanobis', thenDistanceParameteris the covariance matrix in the Mahalanobis distance formula. TheCovname-value argument oflofsets this property.If
Distanceis'minkowski', thenDistanceParameteris the exponent in the Minkowski distance formula. TheExponentname-value argument oflofsets this property.
This property is read-only.
Tie inclusion flag indicating whether LocalOutlierFactor includes
all the neighbors whose distance values are equal to the kth smallest
distance, specified as logical 0 (false) or
1 (true). If IncludeTies is
true, LocalOutlierFactor includes all of these
neighbors. Otherwise, LocalOutlierFactor includes exactly
k neighbors.
This property is read-only.
Number of nearest neighbors in X used to compute local outlier
factor values, specified as a positive integer value.
This property is read-only.
Predictor variable names, returned as a cell array of character vectors. The order of the
elements in PredictorNames corresponds to the order in which the
predictor names appear in the training data.
This property is read-only.
Threshold for the anomaly score used to identify anomalies in the training data, specified as a nonnegative scalar.
The software identifies observations with anomaly scores above the threshold as anomalies.
The
loffunction determines the threshold value to detect the specified fraction (ContaminationFractionproperty) of training observations as anomalies.
The
isanomalyobject function uses theScoreThresholdproperty value as the default value of theScoreThresholdname-value argument.
This property is read-only.
Nearest neighbor search method, specified as 'kdtree' or
'exhaustive'.
'kdtree'— This method uses a Kd-tree algorithm to find nearest neighbors. This option is valid when the distance metric (Distance) is one of the following:'euclidean'— Euclidean distance'cityblock'— City block distance'minkowski'— Minkowski distance'chebychev'— Chebychev distance
'exhaustive'— This method uses the exhaustive search algorithm to find nearest neighbors.When you compute local outlier factor values for
Xusing theloffunction, the function finds nearest neighbors by computing the distance values from all points inXto each point inX.When you compute local outlier factor values for new data
Xnewusing theisanomalyfunction, the function finds nearest neighbors by computing the distance values from all points inXto each point inXnew.
Object Functions
isanomaly | Find anomalies in data using local outlier factor |
Examples
Detect outliers (anomalies in training data) by using the lof function.
Load the sample data set NYCHousing2015.
load NYCHousing2015The data set includes 10 variables with information on the sales of properties in New York City in 2015. Display a summary of the data set.
summary(NYCHousing2015)
NYCHousing2015: 91446×10 table
Variables:
BOROUGH: double
NEIGHBORHOOD: cell array of character vectors
BUILDINGCLASSCATEGORY: cell array of character vectors
RESIDENTIALUNITS: double
COMMERCIALUNITS: double
LANDSQUAREFEET: double
GROSSSQUAREFEET: double
YEARBUILT: double
SALEPRICE: double
SALEDATE: datetime
Statistics for applicable variables:
NumMissing Min Median Max Mean Std
BOROUGH 0 1 3 5 2.8431 1.3343
NEIGHBORHOOD 0
BUILDINGCLASSCATEGORY 0
RESIDENTIALUNITS 0 0 1 8759 2.1789 32.2738
COMMERCIALUNITS 0 0 0 612 0.2201 3.2991
LANDSQUAREFEET 0 0 1700 29305534 2.8752e+03 1.0118e+05
GROSSSQUAREFEET 0 0 1056 8942176 4.6598e+03 4.3098e+04
YEARBUILT 0 0 1939 2016 1.7951e+03 526.9998
SALEPRICE 0 0 333333 4.1111e+09 1.2364e+06 2.0130e+07
SALEDATE 0 01-Jan-2015 09-Jul-2015 31-Dec-2015 07-Jul-2015 2470:47:17
Remove nonnumeric variables from NYCHousing2015. The data type of the BOROUGH variable is double, but it is a categorical variable indicating the borough in which the property is located. Remove the BOROUGH variable as well.
NYCHousing2015 = NYCHousing2015(:,vartype("numeric"));
NYCHousing2015.BOROUGH = [];Train a local outlier factor model for NYCHousing2015. Specify the fraction of anomalies in the training observations as 0.01.
[Mdl,tf,scores] = lof(NYCHousing2015,ContaminationFraction=0.01);
Mdl is a LocalOutlierFactor object. lof also returns the anomaly indicators (tf) and anomaly scores (scores) for the training data NYCHousing2015.
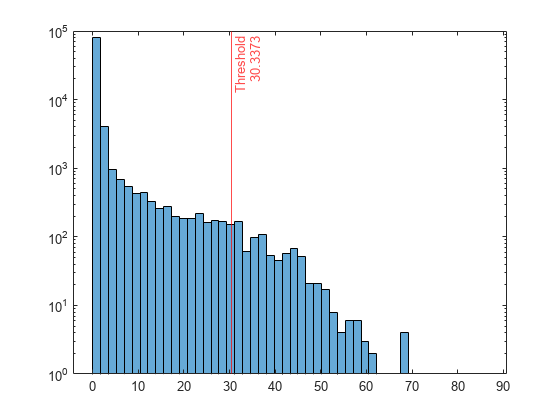
Plot a histogram of the score values. Create a vertical line at the score threshold corresponding to the specified fraction.
h = histogram(scores,NumBins=50); h.Parent.YScale = 'log'; xline(Mdl.ScoreThreshold,"r-",["Threshold" Mdl.ScoreThreshold])
If you want to identify anomalies with a different contamination fraction (for example, 0.05), you can train a new local outlier factor model.
[newMdl,newtf,scores] = lof(NYCHousing2015,ContaminationFraction=0.05);
Note that changing the contamination fraction changes the anomaly indicators only, and does not affect the anomaly scores. Therefore, if you do not want to compute the anomaly scores again by using lof, you can obtain a new anomaly indicator with the existing score values.
Change the fraction of anomalies in the training data to 0.05.
newContaminationFraction = 0.05;
Find a new score threshold by using the quantile function.
newScoreThreshold = quantile(scores,1-newContaminationFraction)
newScoreThreshold = 6.7493
Obtain a new anomaly indicator.
newtf = scores > newScoreThreshold;
Create a LocalOutlierFactor object for uncontaminated training observations by using the lof function. Then detect novelties (anomalies in new data) by passing the object and the new data to the object function isanomaly.
Load the 1994 census data stored in census1994.mat. The data set consists of demographic data from the US Census Bureau to predict whether an individual makes over $50,000 per year.
load census1994census1994 contains the training data set adultdata and the test data set adulttest. The predictor data must be either all continuous or all categorical to train a LocalOutlierFactor object. Remove nonnumeric variables from adultdata and adulttest.
adultdata = adultdata(:,vartype("numeric")); adulttest = adulttest(:,vartype("numeric"));
Train a local outlier factor model for adultdata. Assume that adultdata does not contain outliers.
[Mdl,tf,s] = lof(adultdata);
Mdl is a LocalOutlierFactor object. lof also returns the anomaly indicators tf and anomaly scores s for the training data adultdata. If you do not specify the ContaminationFraction name-value argument as a value greater than 0, then lof treats all training observations as normal observations, meaning all the values in tf are logical 0 (false). The function sets the score threshold to the maximum score value. Display the threshold value.
Mdl.ScoreThreshold
ans = 28.6719
Find anomalies in adulttest by using the trained local outlier factor model.
[tf_test,s_test] = isanomaly(Mdl,adulttest);
The isanomaly function returns the anomaly indicators tf_test and scores s_test for adulttest. By default, isanomaly identifies observations with scores above the threshold (Mdl.ScoreThreshold) as anomalies.
Create histograms for the anomaly scores s and s_test. Create a vertical line at the threshold of the anomaly scores.
h1 = histogram(s,NumBins=50,Normalization="probability"); hold on h2 = histogram(s_test,h1.BinEdges,Normalization="probability"); xline(Mdl.ScoreThreshold,"r-",join(["Threshold" Mdl.ScoreThreshold])) h1.Parent.YScale = 'log'; h2.Parent.YScale = 'log'; legend("Training Data","Test Data",Location="north") hold off

Display the observation index of the anomalies in the test data.
find(tf_test)
ans = 0×1 empty double column vector
The anomaly score distribution of the test data is similar to that of the training data, so isanomaly does not detect any anomalies in the test data with the default threshold value. You can specify a different threshold value by using the ScoreThreshold name-value argument. For an example, see Specify Anomaly Score Threshold.
More About
The local outlier factor (LOF) algorithm detects anomalies based on the relative density of an observation with respect to the surrounding neighborhood.
The algorithm finds the k-nearest neighbors of an observation and computes the local reachability densities for the observation and its neighbors. The local outlier factor is the average density ratio of the observation to its neighbor. That is, the local outlier factor of observation p is
where
lrdk(·) is the local reachability density of an observation.
Nk(p) represents the k-nearest neighbors of observation p. You can specify the
IncludeTiesname-value argument astrueto include all the neighbors whose distance values are equal to the kth smallest distance, or specifyfalseto include exactly k neighbors. The defaultIncludeTiesvalue oflofisfalsefor more efficient performance. Note that the algorithm in [1] uses all the neighbors.|Nk(p)| is the number of observations in Nk(p).
For normal observations, the local outlier factor values are less than or close to 1,
indicating that the local reachability density of an observation is higher than or similar
to its neighbors. A local outlier factor value greater than 1 can indicate an anomaly. The
ContaminationFraction argument of lof and the ScoreThreshold
argument of isanomaly control the threshold for the local outlier
factor values.
The algorithm measures the density based on the reachability distance. The reachability distance of observation p with respect to observation o is defined as
where
dk(o) is the kth smallest distance among the distances from observation o to its neighbors.
d(p,o) is the distance between observation p and observation o.
The algorithm uses the reachability distance to reduce the statistical fluctuations of d(p,o) for the observations close to observation o.
The local reachability density of observation p is the reciprocal of the average reachability distance from observation p to its neighbors.
The density value can be infinity if the number of duplicates is greater than the number of
neighbors (k). Therefore, if the training data contains duplicates, the
lof and isanomaly functions use the weighted
local outlier factor (WLOF) algorithm. This algorithm computes the weighted local outlier
factors using the weighted local reachability density (wlrd).
where
and w(o) is the number of duplicates for observation o in the training data. After computing the weight values, the algorithm treats each set of duplicates as one observation.
A distance metric is a function that defines a distance between two observations. LocalOutlierFactor supports various distance metrics for continuous variables and categorical variables.
Given an mx-by-n data matrix X, which is treated as mx (1-by-n) row vectors x1, x2, ..., xmx, and an my-by-n data matrix Y, which is treated as my (1-by-n) row vectors y1, y2, ...,ymy, the various distances between the vector xs and yt are defined as follows:
Distance metrics for continuous (numeric) variables
Euclidean distance
The Euclidean distance is a special case of the Minkowski distance, where p = 2.
Fast Euclidean distance is the same as Euclidean distance, but uses an algorithm that usually saves time when the number of variables in an observation n exceeds 10. See Algorithms.
Mahalanobis distance
where C is the covariance matrix.
City block distance
The city block distance is a special case of the Minkowski distance, where p = 1.
Minkowski distance
For the special case of p = 1, the Minkowski distance gives the city block distance. For the special case of p = 2, the Minkowski distance gives the Euclidean distance. For the special case of p = ∞, the Minkowski distance gives the Chebychev distance.
Chebychev distance
The Chebychev distance is a special case of the Minkowski distance, where p = ∞.
Cosine distance
Correlation distance
where
and
Spearman distance is one minus the sample Spearman's rank correlation between observations (treated as sequences of values):
where
Distance metrics for categorical variables
Hamming distance
The Hamming distance is the percentage of coordinates that differ.
Jaccard distance is one minus the Jaccard coefficient, which is the percentage of nonzero coordinates that differ:
Tips
You can use interpretability features, such as
lime,shapley,partialDependence, andplotPartialDependence, to interpret how predictors contribute to anomaly scores. Define a custom function that returns anomaly scores, and then pass the custom function to the interpretability functions. For an example, see Specify Model Using Function Handle.
References
[1] Breunig, Markus M., et al. “LOF: Identifying Density-Based Local Outliers.” Proceedings of the 2000 ACM SIGMOD International Conference on Management of Data, 2000, pp. 93–104.
Version History
Introduced in R2022bThe lof function gains support for the
"fasteuclidean"
Distance algorithm. This algorithm usually computes distances faster
than the default "euclidean" algorithm when the number of variables in a
data point exceeds 10. The algorithm, described in Algorithms, uses extra memory to
store an intermediate Gram matrix. Set the size of this memory allocation using the
CacheSize argument.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Sélectionner un site web
Choisissez un site web pour accéder au contenu traduit dans votre langue (lorsqu'il est disponible) et voir les événements et les offres locales. D’après votre position, nous vous recommandons de sélectionner la région suivante : .
Vous pouvez également sélectionner un site web dans la liste suivante :
Comment optimiser les performances du site
Pour optimiser les performances du site, sélectionnez la région Chine (en chinois ou en anglais). Les sites de MathWorks pour les autres pays ne sont pas optimisés pour les visites provenant de votre région.
Amériques
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)