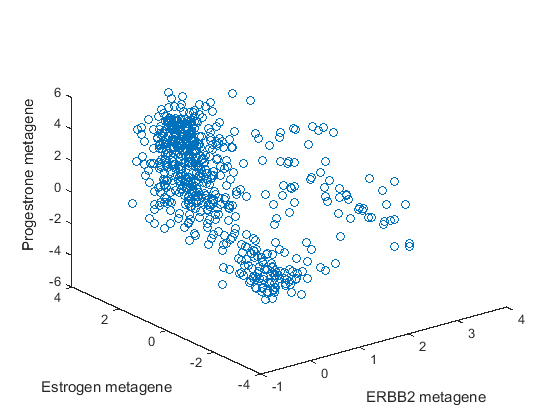
metafeatures
Attractor metagene algorithm for feature engineering using mutual information-based learning
Syntax
Description
M = metafeatures(X)M in X using
the attractor metagene algorithm described in [1].
M is a r-by-n matrix. r is
the number of metafeatures identified during each repetition of the
algorithm. The default number of repetitions is 1. By default, only
unique metafeatures are returned in M. If multiple repetitions result
in the same metafeature, then just one copy is returned in M. n is
the number of samples (patients or time points).
X is a p-by-n numeric
matrix. p is the number of variables, features,
or genes. In other words, rows of X correspond
to variables, such as measurements of gene expression for different
genes. Columns correspond to different samples, such as patients or
time points.
[
uses a p-by-1 cell array of character vectors or string vector
M,W,GSorted]
= metafeatures(X,G)G containing the variable names and returns a
p-by-r cell array of variable names
GSorted sorted by the decreasing weight.
The ith column of GSorted lists
the feature (variable) names in order of their contributions to the ith
metafeature.
[ returns the indices M,W,GSorted,GSortedInd]
= metafeatures(___)GSortedInd such
that GSorted = G(GSortedInd).
[___] = metafeatures(___,
uses additional options specified by one or more Name,Value)Name,Value pair
arguments.
[___] = metafeatures( uses
a p-by-n table T)T.
Gene names are the row names of the table. M = W'*T{:,:}.
[___] = metafeatures(
uses additional options specified by one or more T,Name,Value)Name,Value pair
arguments.
Note
It is possible that the number of metafeatures (r)
returned in M can be fewer than the number of
replicates (repetitions). Even though you may have set the number
of replicates to a positive integer greater than 1, if each repetition
returns the same metafeature, then r is 1, and M is
1-by-n. This is because, by default, the function
returns only unique metafeatures. If you prefer to get all metafeatures,
set 'ReturnUnique' to false.
A metafeature is considered unique if the Pearson correlation between
it and all previously found metafeatures is less than the 'UniqueTolerance' value
(the default value is 0.98).
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
References
[1] Cheng, W-Y., Ou Yang, T-H., and Anastassiou, D. (2013). Biomolecular events in cancer revealed by attractor metagenes. PLoS Computational Biology 9(2): e1002920.
[2] Daub, C., Steuer, R., Selbig, J., and Kloska, S. (2004). Estimating mutual information using B-spline functions – an improved similarity measure for analysing gene expression data. BMC Bioinformatics 5, 118.
[3] Hefti, M.M., Hu, R., Knoblauch, N.W., Collins, L.C., Haibe-Kains, B., Tamimi, R.M., and Beck, A.H. (2013). Estrogen receptor negative/progesterone receptor positive breast cancer is not a reproducible subtype. Breast Cancer Research. 15:R68.