fairnessWeights
Syntax
Description
weights = fairnessWeights(Tbl,AttributeName,ResponseVarName)AttributeName sensitive attribute and
the ResponseVarName response variable in the data set
Tbl. For every combination of a group in the sensitive attribute and
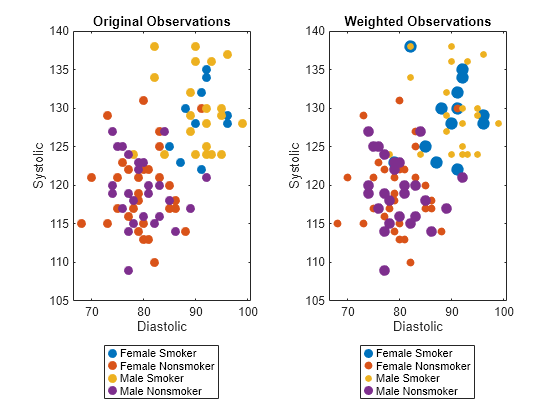
a class label in the response variable, the software computes a weight value. The function
then assigns each observation in Tbl its corresponding weight. The
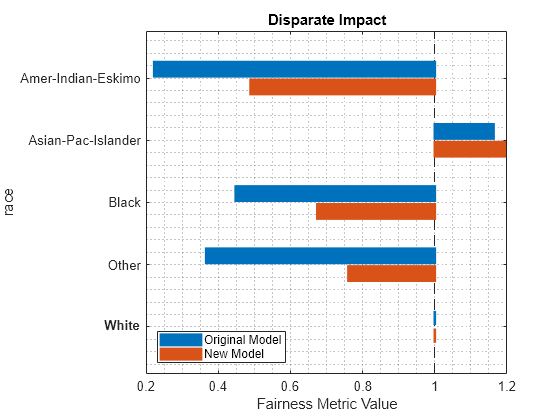
returned weights vector introduces fairness across the sensitive
attribute groups. For more information, see Algorithms.
weights = fairnessWeights(Tbl,AttributeName,Y)Y.
weights = fairnessWeights(___,Weights=initialWeights)initialWeights before
computing the fairness weights, using any of the input argument combinations in previous
syntaxes. These initial weights are typically used to capture some aspect of the data set
that is unrelated to the sensitive attribute, such as expected class distributions.
Examples
Input Arguments
Output Arguments
Algorithms
Assume x is an observation in class k with sensitive attribute
g. If you do not specify initial weights
(initialWeights), then the fairnessWeights
function assigns the following fairness weight to the observation: .
ng is the number of observations with sensitive attribute g.
nk is the number of observations in class k.
ngk is the number of observations in class k with sensitive attribute g.
n is the total number of observations.
is the ideal probability of an observation having sensitive attribute g and being in class k—that is, the product of the probability of an observation having sensitive attribute g and the probability of an observation being in class k. Note that this equation holds for the true probability if the sensitive attribute and the response variable are independent.
is the observed probability of an observation having sensitive attribute g and being in class k.
For more information, see [1].
If you specify initial weights, then the function computes fw(x) using the sum of the initial weights rather than the number of observations. For example, instead of using ng, the function uses the sum of the initial weights of the observations with sensitive attribute g.
References
[1] Kamiran, Faisal, and Toon Calders. “Data Preprocessing Techniques for Classification without Discrimination.” Knowledge and Information Systems 33, no. 1 (October 2012): 1–33. https://doi.org/10.1007/s10115-011-0463-8.
Version History
Introduced in R2022b