RobustRandomCutForest
Description
Use a robust random cut forest model object
RobustRandomCutForest for outlier detection and novelty
detection.
Outlier detection (detecting anomalies in training data) — Detect anomalies in training data by using the
rrcforestfunction. Therrcforestfunction returns aRobustRandomCutForestmodel object, anomaly indicators, and scores for the training data.Novelty detection (detecting anomalies in new data with uncontaminated training data) — Create a
RobustRandomCutForestmodel object by passing uncontaminated training data (data with no outliers) torrcforest. Detect anomalies in new data by passing the object and the new data to the object functionisanomaly. Theisanomalyfunction returns anomaly indicators and scores for the new data.
Creation
Create a RobustRandomCutForest model object by using the rrcforest
function.
Properties
This property is read-only.
Categorical predictor
indices, returned as a vector of positive integers. CategoricalPredictors
contains index values indicating that the corresponding predictors are categorical. The index
values are between 1 and p, where p is the number of
predictors used to train the model. If none of the predictors are categorical, then this
property is empty ([]).
This property is read-only.
Collusive displacement calculation method, specified as 'maximal'
or 'average'.
The software finds the maximum change ('maximal') or the average
change ('average') in model complexity for each tree, and computes
the collusive displacement (anomaly score) for each observation. For details, see Anomaly Scores.
This property is read-only.
Fraction of anomalies in the training data, specified as a numeric scalar in the range [0,1].
If the
ContaminationFractionvalue is 0, thenrrcforesttreats all training observations as normal observations, and sets the score threshold (ScoreThresholdproperty value) to the maximum anomaly score value of the training data.If the
ContaminationFractionvalue is in the range (0,1], thenrrcforestdetermines the threshold value (ScoreThresholdproperty value) so that the function detects the specified fraction of training observations as anomalies.
This property is read-only.
Predictor means of the training data, specified as a numeric vector.
If you specify
StandardizeData=truewhen you train a robust random cut forest model usingrrcforest:The length of
Muis equal to the number of predictors.If you set
StandardizeData=false, thenMuis an empty vector ([]).
This property is read-only.
Number of robust random cut trees (trees in the robust random cut forest model), specified as a positive integer scalar.
This property is read-only.
Number of observations to draw from the training data without replacement for each robust random cut tree (tree in the robust random cut forest model), specified as a positive integer scalar.
This property is read-only.
Predictor variable names, returned as a cell array of character vectors. The order of the
elements in PredictorNames corresponds to the order in which the
predictor names appear in the training data.
This property is read-only.
Threshold for the anomaly score used to identify anomalies in the training data,
specified as a numeric scalar in the range [0,Inf).
The software identifies observations with anomaly scores above the threshold as anomalies.
The
rrcforestfunction determines the threshold value to detect the specified fraction (ContaminationFractionproperty) of training observations as anomalies.
The
isanomalyobject function uses theScoreThresholdproperty value as the default value of theScoreThresholdname-value argument.
This property is read-only.
Predictor standard deviations of the training data, specified as a numeric vector.
If you specify
StandardizeData=truewhen you train a robust random cut forest model usingrrcforest:The length of
Sigmais equal to the number of predictors.If you set
StandardizeData=false, thenSigmais an empty vector ([]).
Object Functions
isanomaly | Find anomalies in data using robust random cut forest |
incrementalLearner | Convert robust random cut forest model to incremental learner |
Examples
Detect outliers (anomalies in training data) by using the rrcforest function.
Load the sample data set NYCHousing2015.
load NYCHousing2015The data set includes 10 variables with information on the sales of properties in New York City in 2015. Display a summary of the data set.
summary(NYCHousing2015)
NYCHousing2015: 91446×10 table
Variables:
BOROUGH: double
NEIGHBORHOOD: cell array of character vectors
BUILDINGCLASSCATEGORY: cell array of character vectors
RESIDENTIALUNITS: double
COMMERCIALUNITS: double
LANDSQUAREFEET: double
GROSSSQUAREFEET: double
YEARBUILT: double
SALEPRICE: double
SALEDATE: datetime
Statistics for applicable variables:
NumMissing Min Median Max Mean Std
BOROUGH 0 1 3 5 2.8431 1.3343
NEIGHBORHOOD 0
BUILDINGCLASSCATEGORY 0
RESIDENTIALUNITS 0 0 1 8759 2.1789 32.2738
COMMERCIALUNITS 0 0 0 612 0.2201 3.2991
LANDSQUAREFEET 0 0 1700 29305534 2.8752e+03 1.0118e+05
GROSSSQUAREFEET 0 0 1056 8942176 4.6598e+03 4.3098e+04
YEARBUILT 0 0 1939 2016 1.7951e+03 526.9998
SALEPRICE 0 0 333333 4.1111e+09 1.2364e+06 2.0130e+07
SALEDATE 0 01-Jan-2015 09-Jul-2015 31-Dec-2015 07-Jul-2015 2470:47:17
The SALEDATE column is a datetime array, which is not supported by rrcforest. Create columns for the month and day numbers of the datetime values, and then delete the SALEDATE column.
[~,NYCHousing2015.MM,NYCHousing2015.DD] = ymd(NYCHousing2015.SALEDATE); NYCHousing2015.SALEDATE = [];
The columns BOROUGH, NEIGHBORHOOD, and BUILDINGCLASSCATEGORY contain categorical predictors. Display the number of categories for the categorical predictors.
length(unique(NYCHousing2015.BOROUGH))
ans = 5
length(unique(NYCHousing2015.NEIGHBORHOOD))
ans = 254
length(unique(NYCHousing2015.BUILDINGCLASSCATEGORY))
ans = 48
For a categorical variable with more than 64 categories, the rrcforest function uses an approximate splitting method that can reduce the accuracy of the robust random cut forest model. Remove the NEIGHBORHOOD column, which contains a categorical variable with 254 categories.
NYCHousing2015.NEIGHBORHOOD = [];
Train a robust random cut forest model for NYCHousing2015. Specify the fraction of anomalies in the training observations as 0.1, and specify the first variable (BOROUGH) as a categorical predictor. The first variable is a numeric array, so rrcforest assumes it is a continuous variable unless you specify the variable as a categorical variable.
rng("default") % For reproducibility [Mdl,tf,scores] = rrcforest(NYCHousing2015, ... ContaminationFraction=0.1,CategoricalPredictors=1);
Mdl is a RobustRandomCutForest model object. rrcforest also returns the anomaly indicators (tf) and anomaly scores (scores) for the training data NYCHousing2015.
Plot a histogram of the score values. Create a vertical line at the score threshold corresponding to the specified fraction.
histogram(scores) xline(Mdl.ScoreThreshold,"r-",["Threshold" Mdl.ScoreThreshold])

If you want to identify anomalies with a different contamination fraction (for example, 0.01), you can train a new robust random cut forest model.
rng("default") % For reproducibility [newMdl,newtf,scores] = rrcforest(NYCHousing2015, ... ContaminationFraction=0.01,CategoricalPredictors=1);
If you want to identify anomalies with a different score threshold value (for example, 65), you can pass the RobustRandomCutForest model object, the training data, and a new threshold value to the isanomaly function.
[newtf,scores] = isanomaly(Mdl,NYCHousing2015,ScoreThreshold=65);
Note that changing the contamination fraction or score threshold changes the anomaly indicators only, and does not affect the anomaly scores. Therefore, if you do not want to compute the anomaly scores again by using rrcforest or isanomaly, you can obtain a new anomaly indicator using the existing score values.
Change the fraction of anomalies in the training data to 0.01.
newContaminationFraction = 0.01;
Find a new score threshold by using the quantile function.
newScoreThreshold = quantile(scores,1-newContaminationFraction)
newScoreThreshold = 63.2642
Obtain a new anomaly indicator.
newtf = scores > newScoreThreshold;
Create a RobustRandomCutForest model object for uncontaminated training observations by using the rrcforest function. Then detect novelties (anomalies in new data) by passing the object and the new data to the object function isanomaly.
Load the 1994 census data stored in census1994.mat. The data set contains demographic data from the US Census Bureau to predict whether an individual makes over $50,000 per year.
load census1994census1994 contains the training data set adultdata and the test data set adulttest.
Assume that adultdata does not contain outliers. Train a robust random cut forest model for adultdata. Specify StandardizeData as true to standardize the input data.
rng("default") % For reproducibility [Mdl,tf,s] = rrcforest(adultdata,StandardizeData=true);
Mdl is a RobustRandomCutForest model object. rrcforest also returns the anomaly indicators tf and anomaly scores s for the training data adultdata. If you do not specify the ContaminationFraction name-value argument as a value greater than 0, then rrcforest treats all training observations as normal observations, meaning all the values in tf are logical 0 (false). The function sets the score threshold to the maximum score value. Display the threshold value.
Mdl.ScoreThreshold
ans = 86.5315
Find anomalies in adulttest by using the trained robust random cut forest model. Because you specified StandardizeData=true when you trained the model, the isanomaly function standardizes the input data by using the predictor means and standard deviations of the training data stored in the Mu and Sigma properties, respectively.
[tf_test,s_test] = isanomaly(Mdl,adulttest);
The isanomaly function returns the anomaly indicators tf_test and scores s_test for adulttest. By default, isanomaly identifies observations with scores above the threshold (Mdl.ScoreThreshold) as anomalies.
Create histograms for the anomaly scores s and s_test. Create a vertical line at the threshold of the anomaly scores.
histogram(s,Normalization="probability") hold on histogram(s_test,Normalization="probability") xline(Mdl.ScoreThreshold,"r-",join(["Threshold" Mdl.ScoreThreshold])) legend("Training Data","Test Data",Location="northwest") hold off

Display the observation index of the anomalies in the test data.
find(tf_test)
ans = 3541
The anomaly score distribution of the test data is similar to that of the training data, so isanomaly detects a small number of anomalies in the test data with the default threshold value.
Zoom in to see the anomaly and the observations near the threshold.
xlim([50 92]) ylim([0 0.001])
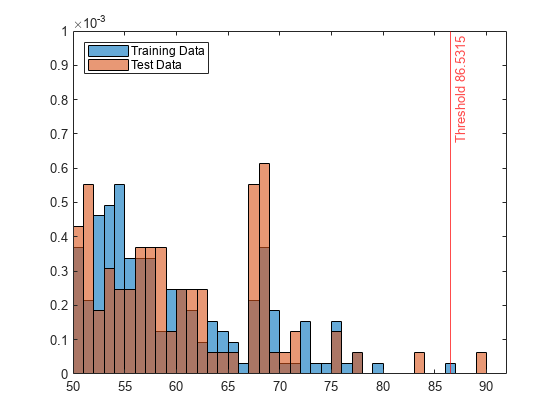
You can specify a different threshold value by using the ScoreThreshold name-value argument. For an example, see Specify Anomaly Score Threshold.
More About
The robust random cut forest algorithm [1] classifies a point as a normal point or an anomaly based on the change in model complexity introduced by the point. Similar to the Isolation Forest algorithm, the robust random cut forest algorithm builds an ensemble of trees. The two algorithms differ in how they choose a split variable in the trees and how they define anomaly scores.
The rrcforest function creates a robust random cut forest model (ensemble
of robust random cut trees) for training observations and detects outliers (anomalies in the
training data). Each tree is trained for a subset of training observations as follows:
rrcforestdraws samples without replacement from the training observations for each tree.rrcforestgrows a tree by choosing a split variable in proportion to the ranges of variables, and choosing the split position uniformly at random. The function continues until every sample reaches a separate leaf node for each tree.
Using the range information in to choose a split variable makes the algorithm robust to irrelevant variables.
Anomalies are easy to describe, but make describing the remainder of the data more
difficult. Therefore, adding an anomaly to a model increases the model complexity of a
forest model [1]. The rrcforest
function identifies outliers using anomaly scores that are defined
based on the change in model complexity.
The isanomaly function uses a trained robust random cut forest model to
detect anomalies in the data. For novelty detection (detecting anomalies in new data with
uncontaminated training data), you can train a robust random cut forest model with
uncontaminated training data (data with no outliers) and use it to detect anomalies in new
data. For each observation of the new data, the function finds the corresponding leaf node
in each tree, computes the change in model complexity introduced by the leaf nodes, and
returns an anomaly indicator and score.
The robust random cut forest algorithm uses collusive displacement as an anomaly score. The collusive displacement of a point x indicates the contribution of x to the model complexity of a forest model. A small positive anomaly score value indicates a normal observation, and a large positive value indicates an anomaly.
As defined in [1], the model complexity |M(T)| of a tree T is the sum of path lengths (the distance from the root node to the leaf nodes) over all points in the training data Z.
where f(y,Z,T) is the depth of y in tree T. The displacement of x is defined to indicate the expected changes in the model complexity introduced by x.
where T' is a tree over Z – {x}. Disp(x,Z) is the expected number of points in the sibling node of the leaf node
containing x. This definition is not robust to duplicates or
near-duplicates, and can cause outlier masking. To avoid outlier masking, the robust random
cut forest algorithm uses the collusive displacement CoDisp, where a set
C includes x and the colluders of
x.
where T" is a tree over Z – C, and |C| is the number of points in the subtree of T for C.
The default value for the CollusiveDisplacement name-value argument of rrcforest
is "maximal". For each tree, by default, the software finds the set
C that maximizes the ratio Disp(x,C)/|C| by traversing from the leaf node of x to the root node,
as described in [2]. If you specify
CollusiveDisplacement="average"
Tips
You can use interpretability features, such as
lime,shapley,partialDependence, andplotPartialDependence, to interpret how predictors contribute to anomaly scores. Define a custom function that returns anomaly scores, and then pass the custom function to the interpretability functions. For an example, see Specify Model Using Function Handle.
References
[1] Guha, Sudipto, N. Mishra, G. Roy, and O. Schrijvers. "Robust Random Cut Forest Based Anomaly Detection on Streams," Proceedings of The 33rd International Conference on Machine Learning 48 (June 2016): 2712–21.
[2] Bartos, Matthew D., A. Mullapudi, and S. C. Troutman. "rrcf: Implementation of the Robust Random Cut Forest Algorithm for Anomaly Detection on Streams." Journal of Open Source Software 4, no. 35 (2019): 1336.
Version History
Introduced in R2023a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Sélectionner un site web
Choisissez un site web pour accéder au contenu traduit dans votre langue (lorsqu'il est disponible) et voir les événements et les offres locales. D’après votre position, nous vous recommandons de sélectionner la région suivante : .
Vous pouvez également sélectionner un site web dans la liste suivante :
Comment optimiser les performances du site
Pour optimiser les performances du site, sélectionnez la région Chine (en chinois ou en anglais). Les sites de MathWorks pour les autres pays ne sont pas optimisés pour les visites provenant de votre région.
Amériques
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)